Contributing (projects & manual)
This chapter mirrors Lineamientos para colaboradores; figures are reproduced from that repo inside this manual.
Do not upload identifiable data or secrets. Heavy tables stay outside GitHub (e.g. Dropbox, Google Drive) and are linked in the README.
Creating repositories
Step 1 — create the repo
Step 2 — configure metadata & base files
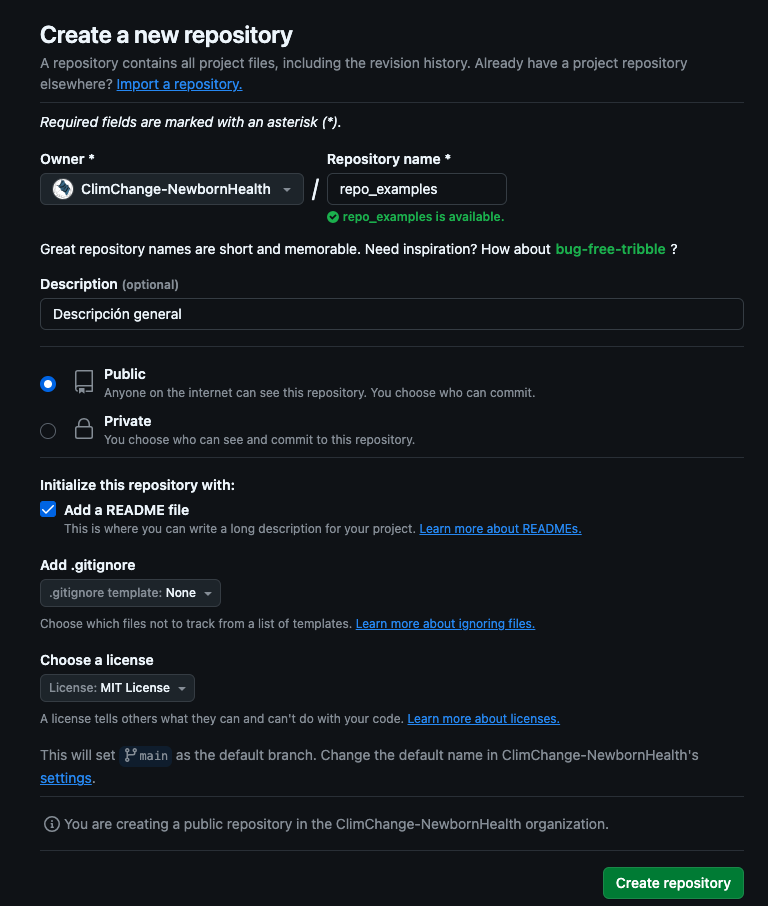
Repositories in the EB-Lab ecosystem under ClimChange-NewbornHealth should have:
- Short names, ideally English.
- A concise description.
- Public visibility (unless a documented exception).
README.md,.gitignore, license (MIT recommended).
Step 3 — clone locally
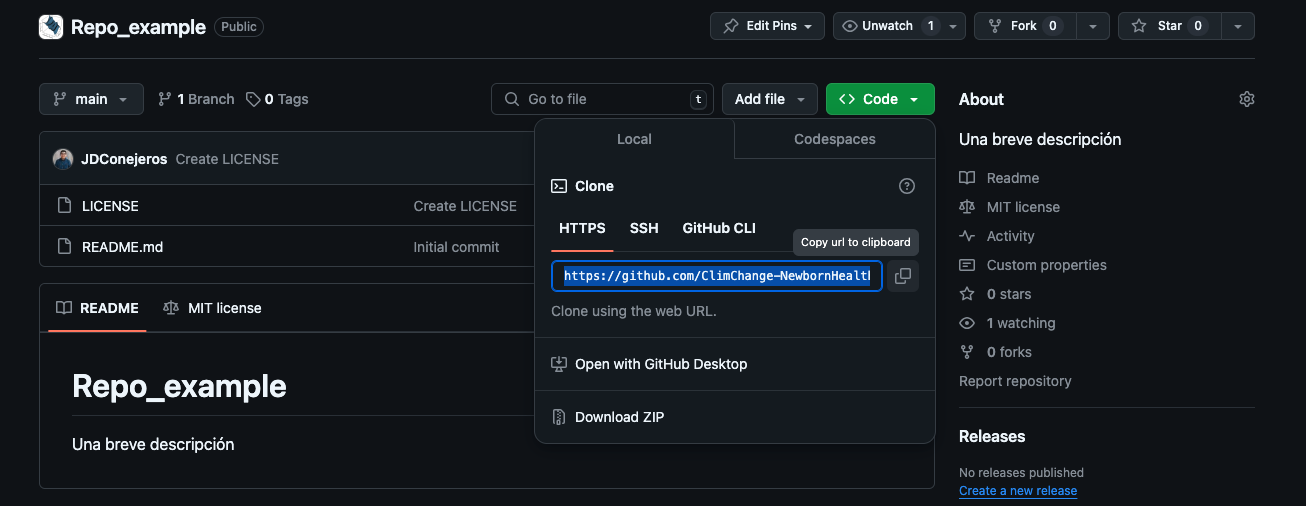
Use RStudio, VS Code, Positron, or any IDE that respects git + relative paths.
Prefer short branches & reviewed PRs before merging to main, especially for Models/ and final artifacts.
Expected folder layout
Code/
Process/
Descriptive/
Models/
Output_Analysis/
Graphs/
Tables/
.gitignore
README.md
LICENSEREADME content
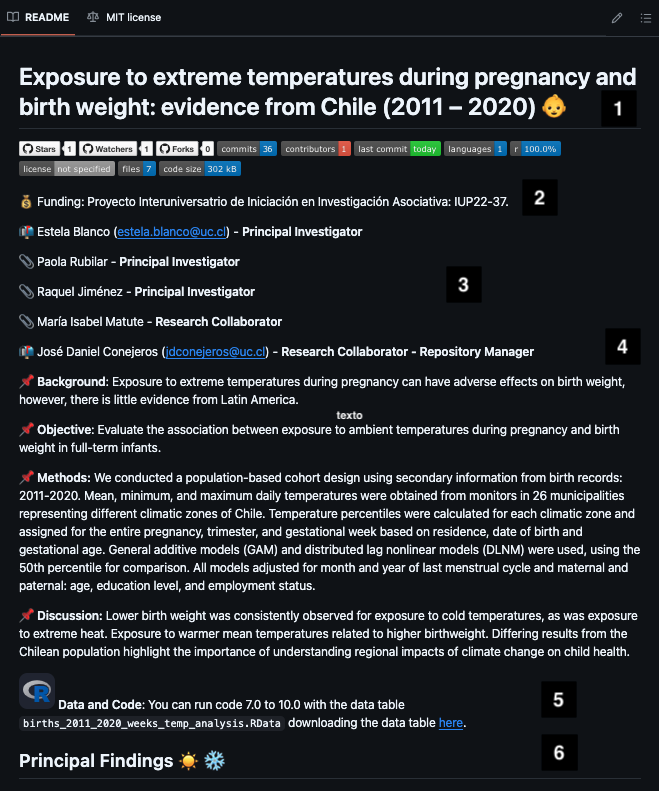
- Title (ideally manuscript title).
- Metrics & funding (see template).
- Authors incl. emails + repo maintainer.
- Scientific description from the abstract where possible.
- Data access links + ordered script execution.
- Key results — numbered figures/tables with notes.
Template: Plantilla_README.md.
Data stays off-repo
Repos hold code to replicate; large tables are linked externally.
Uploading entire bases bloats the repo and increases disclosure risk. Keep access-controlled storage outside public git.

Example README + external data workflow: CIIIA-ClimateBirthWeightAnalysis.
Contributing to this manual
- Fork or obtain write access to
eblab-manual. - Edit
.qmdfiles /_quarto.ymlas needed. - Maintain Spanish + English: run renders for both locales before PRs (
quarto render+cd english && quarto render). - Open a pull request explaining the change.
Repository access questions: jdconejeros@uc.cl (Research collaborator – Repository manager).